 George Hripcsak, MD, MS
George Hripcsak, MD, MS
Vivian Beaumont Allen Professor of Biomedical Informatics, Columbia University
Chair, Department of Biomedical Informatics,
Columbia University
Director, Medical Informatics Services, NewYork-Presbyterian Hospital/Columbia
Biography
George Hripcsak,
MD, MS, is Vivian Beaumont Allen Professor and Chair of Columbia University’s
Department of Biomedical Informatics and Director of Medical Informatics
Services for NewYork-Presbyterian Hospital/Columbia Campus. He is a
board-certified internist with degrees in chemistry, medicine, and
biostatistics. Dr. Hripcsak’s current research focus is on the clinical
information stored in electronic health records and on the development of
next-generation health record systems. Using nonlinear time series analysis,
machine learning, knowledge engineering, and natural language processing, he is
developing the methods necessary to support clinical research and patient
safety initiatives. He leads the Observational Health Data Sciences and
Informatics (OHDSI) coordinating center; OHDSI is an international network with
180 researchers and 600 million patient records. For his work in precision
medicine, he serves as a PI on Columbia’s eMERGE grant, as a PI on Columbia’s
regional recruitment center for the All of Us precision medicine program, and
as site PI for Columbia’s role on the All of Us Data and Research Center. He
co-chaired the Meaningful Use Workgroup of U.S. Department of Health and Human
Services’s Office of the National Coordinator of Health Information Technology;
it defines the criteria by which health care providers collect incentives for
using electronic health records. He led the effort to create the Arden Syntax,
a language for representing health knowledge that has become a national
standard. Dr. Hripcsak is a fellow of the National Academy of Medicine, the
American College of Medical Informatics, and the New York Academy of Medicine,
and he chaired the U.S. National Library of Medicine’s Biomedical Library and
Informatics Review Committee. He has published over 350 papers.
Research
Research openings: I welcome PhD and MD postdoctoral candidates
interested in working on any of the research listed below. Experience in a
relevant methodological area (e.g. nonlinear dynamics) and domain area
(medicine, physiology) would be ideal. I also welcome Columbia graduate and
undergraduate students.
1.
Phenotyping. My work in decision support led me to
realize that the main obstacle was our inability to exploit clinical data
effectively. I therefore helped to create the informatics field now known as
“phenotyping” with early publications in use of symbolic reasoning and machine
learning to map raw health record data to clinical concepts. My work continues
in this area to this day.
a.
Hripcsak G, Johnson SB, Clayton PD. Desperately
seeking data: knowledge base-database links. Proc Annu Symp Comput Appl Med
Care 1993:639-43.
b.
Wilcox
A, Hripcsak G. Knowledge discovery
and data mining to assist natural language understanding. Proc Amia Symp
1998:835-9.
c.
Wilcox
AB, Hripcsak G. The role of domain
knowledge in automating medical text report classification. J Am Med Inform
Assoc 2003;10:330-8. PMC181983
d.
Hripcsak G, Albers DJ. Next-generation phenotyping of
electronic health records. J Am Med Inform Assoc 2013;20:117–21:
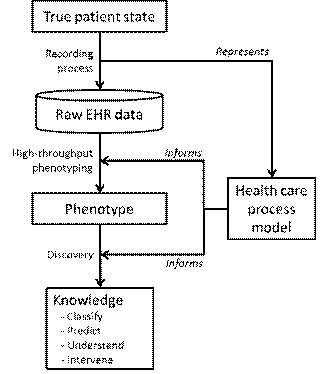
2.
Use of non-linear time series analysis in
clinical phenotyping. My
recent work in phenotyping has turned to the use of non-linear time series
analysis to improve use of clinical data and understanding of physiologic
processes, and I have begun to study the health care process itself and how it
affects the recording of data so that we can reduce the bias associated with
observational studies. This time series work was published not just in the top
informatics journals, but also in top physics journals in non-linear science.
a.
Albers
DJ, Hripcsak G. A statistical
dynamics approach to the study of human health data: resolving population scale
diurnal variation in laboratory data. Physics Letters A 2010;374:1159-64.
b.
Albers
DJ, Hripcsak G. Using time-delayed
mutual information to discover and interpret temporal correlation structure in
complex populations. Chaos 2012;22:013111; doi:10.1063/1.3675621.
c.
Hripcsak G, Albers DJ, Perotte A. Exploiting time in
electronic health record correlations. J Am Med Inform Assoc 2011;18:Suppl 1
i109-i115.
d.
Hripcsak G, Albers DJ, Perotte A. Parameterizing time
in electronic health record studies. J Am Med Inform Assoc 2015 Feb 26. pii:
ocu051. doi: 10.1093/jamia/ocu051.
e.
Albers
DJ, Levine M, Gluckman B, Ginsberg H, Hripcsak
G, Mamykina L. Personalized glucose forecasting for type 2 diabetes using
data assimilation. PLoS Comput Biol 2017;13:e1005232. doi:
10.1371/journal.pcbi.1005232:
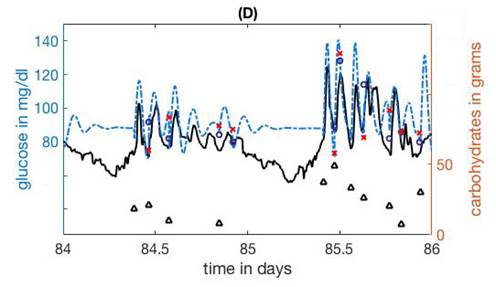
3.
Observational research and bridging
phenotype and genotype. As
PI of the Observational Health Data Sciences and Informatics (OHDSI)
coordinating center, I assist the research community with its 600-million
patient-record network, developing and applying new methods in observational
research. As a PI on the Columbia eMERGE grant, as a PI of the Columbia PMI
Cohort Program regional healthcare provider organization, and as site PI for
Columbia’s role on the PMI Cohort Program Data and Research Support Center, I
work in converting electronic health record data to a form useful for
correlating with genomic information.
a.
Ryan
PB, Madigan D, Stang PE, Schuemie MJ, Hripcsak
G. Medication-wide association studies. CPT: Pharmacometrics & Systems
Pharmacology 2013;2,e76;doi:10.1038/psp.2013.52. PMC4026636
b.
Overby
CL, Pathak J, Gottesman O, Haerian K, Perotte A, Murphy S, Bruce K, Johnson S,
Talwalkar J, Shen Y, Ellis S, Kullo I, Chute C, Friedman C, Bottinger E, Hripcsak G, Weng C. A collaborative
approach to developing an electronic health record phenotyping algorithm for
drug-induced liver injury. J Am Med Inform Assoc 2013 Dec;20(e2):e243-52. doi:
10.1136/amiajnl-2013-001930. PMC3861914
c.
Duke
JD, Ryan PB, Suchard MA, Hripcsak G,
Jin P, Reich C, Schwalm MS, Khoma Y, Wu Y, Xu H, Shah NH, Banda JM, Schuemie
MJ. Risk of angioedema associated with levetiracetam compared with phenytoin:
Findings of the observational health data sciences and informatics research
network. Epilepsia 2017 doi: 10.1111/epi.13828.
d.
Hripcsak G, Ryan P, Duke J, Shah NH, Park RW, Huser
V, Suchard MA, Schuemie M, DeFalco F, Perotte A, Banda J, Reich C, Shilling L,
Matheny M, Meeker D, Pratt N, Madigan D. Addressing clinical questions at
scale: OHDSI characterization of treatment pathways. Proc Natl Acad Sci USA
2016. DOI: 10.1073/pnas.1510502113:
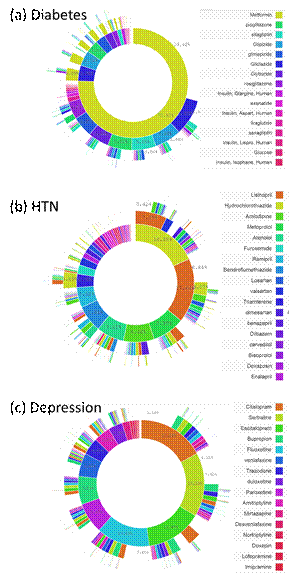
4.
Electronic health records and automated
decision support. My
earliest work was in health knowledge representation and decision support. I
led the creation of the Arden Syntax, which is a standard for representing
health knowledge related to alerts and reminders. It was adopted as an
international standard and today sits under the Health Level Seven (HL7)
standards organization. Paralleling my career-long work building and running
electronic health records and setting national policy for health records
(co-chairing the Meaningful Use Workgroup), I have been publishing on the use
and effect of health records.
a.
Hripcsak G, Hripcsak G, Ludemann P, Pryor TA, Wigertz
OB, Clayton PD. Rationale for the Arden Syntax. Comput Biomed Res
1994;27:291–324.
b.
Shea
S, Hripcsak G. Accelerating the use
of electronic health records in physician practices. NEJM 2010;362:192-5.
c.
Hripcsak G, Vawdrey DK, Fred MR, Bostwick SB. Use of
electronic clinical documentation: time spent and team interactions. J Am Med
Inform Assoc 2011;18:112-7.
d.
Green
RA, Hripcsak G, Salmasian H, Lazar
EJ, Bostwick SB, Bakken SR,Vawdrey DK. Intercepting Wrong-Patient Orders in a
Computerized Provider Order Entry System. Ann Emerg Med 2014 Dec 17. pii:
S0196-0644(14)01558-3. doi: 10.1016/j.annemergmed.2014.11.017:
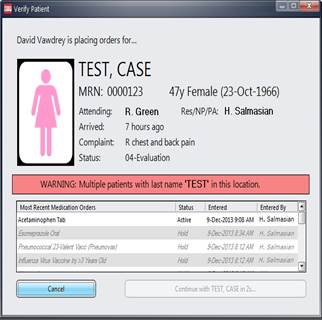
5.
Natural language processing evaluation and
temporal processing. I
developed new methods for evaluating natural language processing and published
the first large-scale evaluation of natural language processing in health care,
followed by the first prospective trial demonstrating that natural language
processing could improve health care quality (respiratory isolation). I
subsequently published studies showing use of natural language processing for
quality measurement and for public health surveillance. I developed new methods
for temporal data in narrative reports with a demonstration of its performance.
a.
Hripcsak G, Friedman C, Alderson PO, DuMouchel W,
Johnson SB, Clayton PD. Unlocking clinical data from narrative reports: a study
of natural language processing. Ann Intern Med 1995;122:681–8.
b.
Knirsch
C, Jain NL, Pablos-Mendez A, Friedman C, Hripcsak
G. Respiratory isolation of tuberculosis patients using clinical guidelines
and an automated clinical decision support system. Infect Control Hosp
Epidemiol 1998;19:94–100.
c.
Hripcsak G, Zhou L, Parsons S, Das AK, Johnson SB.
Modeling electronic discharge summaries as a simple temporal constraint
satisfaction problem. J Am Med Inform Assoc 2005;12:55–63.
d.
Zhou
L, Parsons S, Hripcsak G. The
evaluation of a temporal reasoning system in processing clinical discharge
summaries. J Am Med Inform Assoc 2008;15:99-106. PMC2274869
e.
Hripcsak G, Elhadad N, Chen C, Zhou L, Morrison FP.
Using empirical semantic correlation to interpret temporal assertions in
clinical texts. J Am Med Inform Assoc 2009;16:220-7:
Service
As Director of Medical Informatics Services
for NewYork-Presbyterian Hospital/Columbia, I oversee the clinical data
warehouse, terminology, iNYP, immunization, and physician outreach and
collaborate on clinician documentation, health information exchange, and
patient portals.
Education
We
offer programs at all levels of informatics training, including PhDs, master's
degrees, postdoctoral fellowship, certificate training, and education for
students in medicine, nursing, dentistry, and public health. See www.dbmi.columbia.edu. I am accepting
Columbia graduate and undergraduate students, and I have slots for postdocs in
my areas of research.
Further information
Publications
via Pubmed
Contact:
George
Hripcsak, MD, MS
622 West 168th Street, PH20
New York, NY 10032
Email:
hripcsak[at]columbia.edu
